
These methods are limited as they cannot identify physical associations between IRE1 and its putative substrates, nor they can dissociate direct and indirect effects emanating from IRE1’s nuclease activity. Previous work aimed at identifying IRE1 substrate recognition relied on transcriptomics ( Han et al., 2009 Hollien et al., 2009 Hollien and Weissman, 2006 So et al., 2012). Since the physiological consequences of IRE1 activation are the product of cleavage of ER-targeted transcripts, there is an inherent need to address the issue of substrate selection. 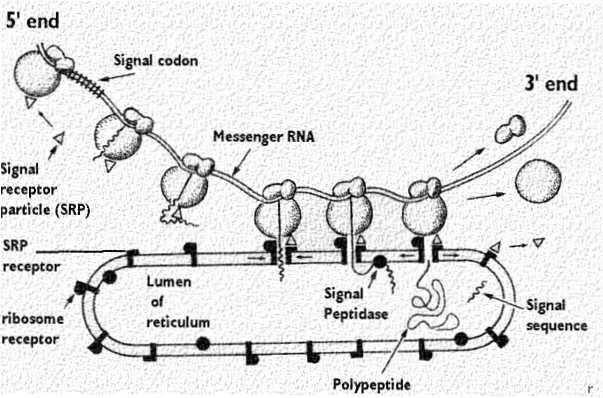
Moreover, the precise determinants controlling IRE1’s exquisite specificity remain limited to our understanding of XBP1 mRNA and a small number of canonical RIDD substrates. 
Despite the progress made in unraveling the ER targeting mechanism of the XBP1 mRNA, our current understanding of recruitment of IRE1 to membrane-bound XBP1 mRNA or RIDD substrate mRNAs is still a mystery. The engagement of the translating XBP1u protein with SRP results in its proper sorting to the ER membrane where it meets IRE1. Peptide sequences encoded in the XBP1u mRNA are recognized by the signal recognition particle (SRP)-dependent co-translational targeting machinery, which is in charge of sorting of most of the mRNAs encoding ER resident, secretory and transmembrane proteins ( Plumb et al., 2015). In metazoans, the XBP1 mRNA is brought to the ER membrane as part of the ribosome-nascent chain complex actively translating XBP1u (‘u’ for unspliced) ( Yanagitani et al., 2009 Yanagitani et al., 2011). In both cases, IRE1 substrate RNAs must be properly targeted to the ER membrane to meet the IRE1 enzyme. Active IRE1 responds to ER stress in two ways: (i) it cleaves an unconventional intron from the mRNA encoding the transcription factor XBP1 (X-box binding protein 1), initiating a spliceosome-independent mRNA splicing reaction that culminates in the production of XBP1s (‘s’ for spliced), a potent transcription activator that increases the ER folding and degradation capacities ( Acosta-Alvear et al., 2007 Reimold et al., 2000 Reimold et al., 2001 Yoshida et al., 1998), and (ii) it cleaves ER-targeted mRNAs in a process known as RIDD (regulated IRE1-dependent decay), thus lowering the ER protein-folding load ( Hollien et al., 2009 Hollien and Weissman, 2006). Active IRE1 molecules trans-autophosphorylate, allowing the subsequent allosteric activation of its C-terminal endoribonuclease domain ( Karagöz et al., 2017 Korennykh et al., 2011). Unfolded proteins serve as direct ligands for IRE1’s lumenal sensor domain, promoting its oligomerization and activation in the plane of the ER membrane ( Aragón et al., 2009 Gardner and Walter, 2011 Karagöz et al., 2017 Kimata et al., 2007 Li et al., 2010). IRE1 is the most evolutionarily conserved sensor and is found from yeast to metazoans. Three ER membrane embedded protein folding sensors control the UPR: ATF6 (activating transcription factor 6), PERK (protein kinase R (PKR)-like kinase) and IRE1 (inositol requiring enzyme 1). The UPR maintains the health of the secreted and membrane-embedded proteome through (i) decreasing ER client protein load, (ii) upregulating chaperones and enzymes that assist protein folding, and (iii) promoting the degradation of misfolded proteins. Perturbations leading to protein folding defects in the ER –collectively known as ER stress– activate an ensemble of transcriptional programs known as the unfolded protein response (UPR)( Karagoz and Acosta-Alvear D, 2018 Walter and Ron, 2011). Protein folding and maturation in the endoplasmic reticulum (ER) are essential for cell physiology, as most of the all secreted and transmembrane proteins are synthesized and folded in this organelle.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
